


Up: User Manual for EGO
Previous: Molecular Dynamics
- 1
-
A. Ansari, J. Berendzen, D. Braunstein, B. R. Cowen, H. Frauenfelder, M. K.
Hong, I. E. T. Iben, J. B. Johnson, P. Ormos, T. B. Sauke, R. Scholl,
A. Schulte, P. J. Steinbach, J. Vittitow, and R. D. Young: ``Rebinding and
relaxation in the myoglobin pocket'',
Biophys. Chem., 26, 337-355 (1987).
- 2
-
H. J. C. Berendsen, J. P. M. Postma, W. F. van Gunsteren, A. DiNola, and J. R.
Haak: ``Molecular dynamics with coupling to an external bath'',
J. Chem. Phys., 81, 3684-3690 (1984).
- 3
-
B. R. Brooks, R. E. Bruccoleri, B. D. Olafson, D. J. States, S. Swaminathan,
and M. Karplus: ``CHARMM: A Program for Macromolecular Energy,
Minimization, and Dynamics Calculations'',
J. Comp. Chem., 4(2), 187-217 (1983).
- 4
-
B. R. Brooks and M. Hodo
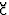 ek: ``Parallelization of CHARMM for MIMD
Machines'',
ek: ``Parallelization of CHARMM for MIMD
Machines'',
Chemical Design Automation News (CDA News), 7(12), 16-22
(1992).
- 5
-
C. L. Brooks III and M. Karplus: ``Deformable stochastic boundaries in
molecular dynamics'',
J. Chem. Phys., 79, 6312-6325 (1983).
- 6
-
A. T. Brünger: ``X-PLOR, Version 2.1'',
The Howard Hughes Medical Institute and Department of Molecular
Biophysics and Biochemistry, Yale University, 260 Whitney Avenue, P.O.
Box 6666, New Haven, CT 06511, 1990.
- 7
-
C. Eckart: ``Some Studies Concerning Rotating Axes and Polyatomic
Molecules'',
Phys. Rev., 47, 552-558 (1935).
- 8
-
N. Ehrenhofer: ``Untersuchung der Konformationsdynamik eines vereinfachten
Proteinmodells'',
Master's thesis, Ludwig-Maximilians-Universität,
München, Germany, 1994.
- 9
-
M. Eichinger: ``Paralleler schneller Multipolalgorithmus mit
Mehrschrittverfahren für Molekulardynamiksimulationen'',
Master's thesis, Ludwig-Maximilians-Universität München,
1995.
- 10
-
M. Eichinger, H. Heller, and H. Grubmüller: ``EGO - An Efficient
Molecular Dynamics Program and its Application to Protein Dynamics
Simulations'',
in R. Esser, P. Grassberger, J. Grotendorst, and M. Lewerenz,
editors, ``Workshop on Molecular Dynamics on Parallel Computers,
John von Neumann Institute for Computing (NIC) Research Centre
Jülich, Germany, 8-10 February 1999'', pages 154-174, Singapore
912805, 2000. World Scientific.
- 11
-
R. Elber and M. Karplus: ``Multiple Conformational States of Proteins: A
Molecular Dynamics Analysis of Myoglobin'',
Science, 235, 318-321 (1987).
- 12
-
M. P. I. Forum: ``MPI: A Message-Passing Interface Standard'',
University of Tennesse, Knoxville, Tennessee 37831, May 1994.
- 13
-
H. Frauenfelder, G. A. Petsko, and D. Tsernoglou: ``Temperature-dependent
X-ray diffraction as a probe of protein structural dynamics'',
Nature (London), 280, 558-563 (1979).
- 14
-
H. Frauenfelder, S. G. Sligar, and P. G. Wolynes: ``The Energy Landscape and
Motions of Proteins'',
Science, 254, 1598-1603 (1991).
- 15
-
A. E. García: ``Large-Amplitude Nonlinear Motions in Proteins'',
Phys. Rev. Lett., 68, 2696-2699 (1992).
- 16
-
C. W. Gardiner: ``Handbook of Stochastic Methods''. Springer-Verlag, Berlin,
Heidelberg, New York, 1985.
- 17
-
A. Geist, A. Beguelin, J. Dongarra, W. Jiang, R. Manchek, and V. Sunderam:
``PVM 3 user's guide and reference manual'',
Oak Ridge National Laboratory, Tennessee 37831, May 1994.
- 18
-
N. G
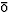 and T. Noguti: ``Structural Basis of Hierarchical Multiple Substates
of a Protein'',
and T. Noguti: ``Structural Basis of Hierarchical Multiple Substates
of a Protein'',
Chem. Scr., 29A, 151-164 (1989).
- 19
-
L. Greengard and V. Rokhlin: ``A Fast Algorithm for Particle Simulations'',
J. Comp. Phys., 73, 325-348 (1987).
- 20
-
L. Greengard and V. Rokhlin: ``On the Evaluation of Electrostatic Interactions
in Molecular Modeling'',
Chem. Scr., 29A, 139-144 (1989).
- 21
-
H. Grubmüller: ``Molekulardynamik von Proteinen auf langen Zeitskalen'',
PhD thesis, Technische Universität München, Germany, Jan. 1994.
- 22
-
H. Grubmüller: ``Predicting slow structural transitions in macromolecular
systems: conformational Flooding'',
Phys. Rev. E, 52, 2893 (1995).
- 23
-
H. Grubmüller, H. Heller, A. Windemuth, and K. Schulten: ``Generalized
Verlet Algorithm for Efficient Molecular Dynamics Simulations with
Long-Range Interactions'',
Mol. Sim., 6, 121-142 (1991).
- 24
-
H. Grubmüller, B. Heymann, and P. Tavan: ``Ligand Binding: Molecular
Mechanics Calculation of the Streptavidin-Biotin Rupture Force'',
Science, 271(5251), 997-999 (1996).
- 25
-
H. Grubmüller and P. Tavan: ``Molecular Dynamics of Conformational
Substates for a Simplified Protein Model'',
J. Chem. Phys., 101, 5047-5057 (1994).
- 26
-
H. Heller: ``Simulation einer Lipidmembran auf einem Parallelrechner'',
Doctoral Dissertation, Technical University of Munich, Germany,
December 1993.
- 27
-
B. Heymann: ``Beschreibung der Streptavidin-Biotin-Bindung mit Hilfe
von Molekulardynamiksimulationen'',
Master's thesis, Ludwig-Maximilians-Universität
München, 1996.
- 28
-
B. W. Kernighan and D. M. Ritchie: ``The C programming language''. Prentice
Hall Software Series, London, 1988.
- 29
-
J. F. Leathrum and J. A. Board: ``The Parallel Fast Multipole Algorithm in
Three Dimensions'',
Technical report, Dept. of Electrical Engineering, Duke University,
Durham, 1992.
- 30
-
M. Levitt and S. Lifson: ``Refinement of Protein Conformation using a
Macromolecular Energy Minimization Procedure'',
J. Mol. Biol., 46, 269-279 (1969).
- 31
-
T. M. Martinetz and K. J. Schulten: ``A `Neural Gas' Network Learns
Topologies.'',
in O. Simula, editor, ``Proceedings of the Int. Conf. on Artificial
Neural Networks, ICANN-91, Espoo, Finland, 24-28 June 1991'', pages
397-402, Amsterdam, 1991. Elsevier Science Publishers.
- 32
-
J. A. McCammon, B. R. Gelin, and M. Karplus: ``Dynamics of Folded
Proteins'',
Nature (London), 267, 585-590 (1977).
- 33
-
C. Niedermeier and P. Tavan: ``A Structure Adapted Multipole Method for
Electrostatic Interactions in Protein Dynamics'',
J. Chem. Phys., 101, 734-748 (1994).
- 34
-
PARSYTEC Computer GmbH, Germany: ``PARIX Version 1.3-PPC reference
manual'', December 1994.
- 35
-
Polygen Corporation, 200 Fifth Av., Waltham, MA 02254, U.S.A.: ``Quanta'',
1988.
- 36
-
E. Shakhnovich, G. Farztdinov, A. M. Gutin, and M. Karplus: ``Protein Folding
Bottlenecks: A Lattice Monte Carlo Simulation'',
Phys. Rev. Lett., 67, 1665-1668 (1991).
- 37
-
W. B. Streett, D. J. Tildesley, and G. Saville: ``Multiple time step methods
in molecular dynamics'',
Mol. Phys., 35, 639-648 (1978).
- 38
-
J. P. Valleau and G. M. Torrie,
in B. J. Berne, editor, ``Statistical Mechanics Part A: Equilibrium
Techniques'', page 137. Plenum Press, New York, 1977.
- 39
-
W. F. van Gunsteren and H. J. C. Berendsen: ``GROMOS Manual'',
BIOMOS b.v., Biomolecular Software, Groningen, The Netherlands.
- 40
-
W. F. van Gunsteren and H. J. C. Berendsen: ``Algorithms for macromolecular
dynamics and constraint dynamics'',
Mol. Phys., 34(5), 1311-1327 (1977).
- 41
-
L. Verlet: ``Computer ``Experiments'' on Classical Fluids.
I. Thermodynamical Properties of Lennard-Jones Molecules'',
Phys. Rev., 159(1), 98-103 (1967).
- 42
-
A. Wallquist and G. Karlström: ``A New Non-Empirical Force Field for
Computer Simulations'',
Chem. Scr., 29A (1989).
- 43
-
S. J. Weiner, P. A. Kollman, D. A. Case, U. C. Singh, C. Ghio, G. Alagona,
J. S. Profeta, and P. Weiner: ``A New Force Field for Molecular Mechanical
Simulation of Nucleic Acids and Proteins'',
J. Am. Chem. Soc., 106, 765-784 (1984).
- 44
-
A. Windemuth: ``Dynamiksimulation von Makromolekülen'',
Diploma Thesis, Technical University of Munich, Physics
Department, T 30, James-Franck-Street, D-8046 Garching / Munich, August 1988.
Helmut Heller
2000-04-19